2. Gallery¶
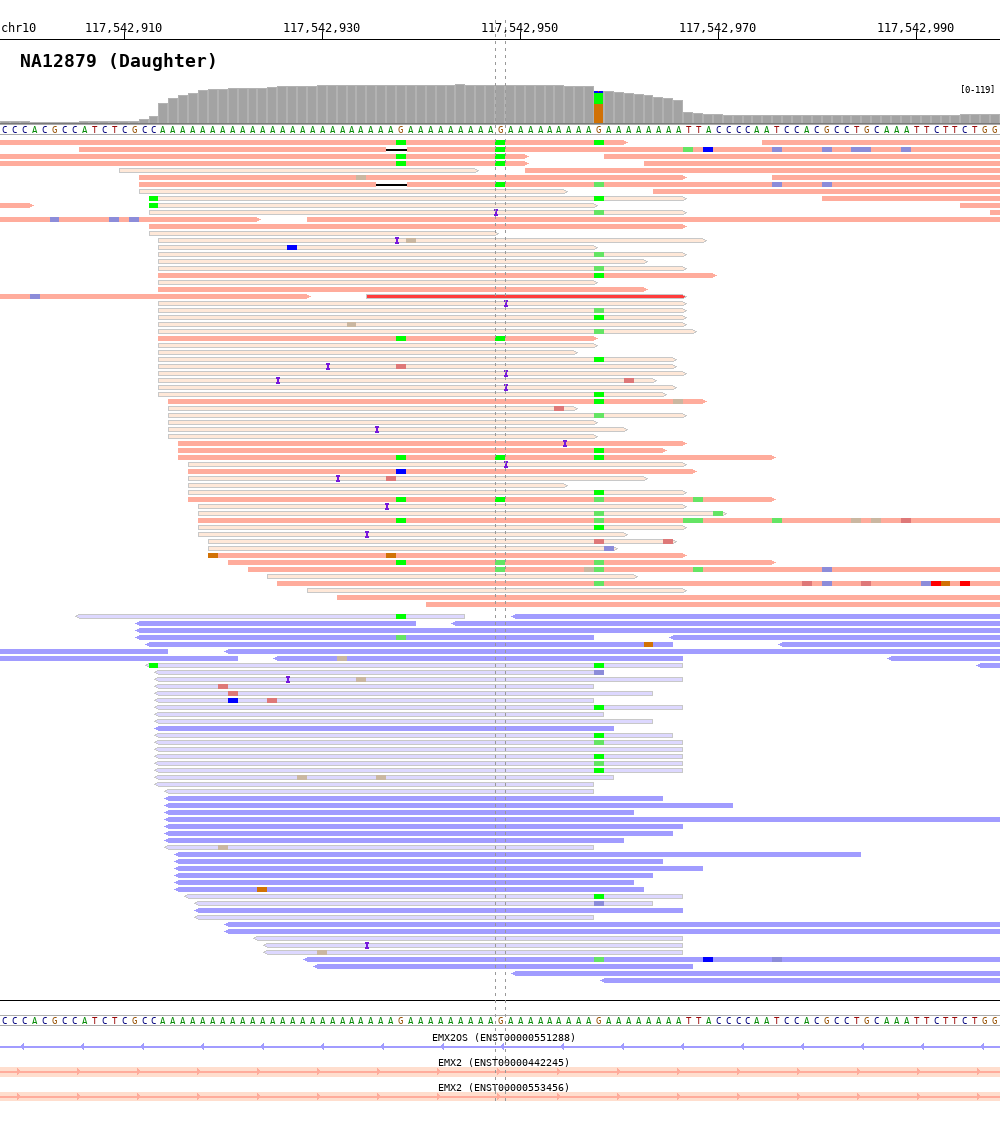
$ bamsnap \
-bam ./data/NA12879.bam \
-title "NA12879 (Daughter)" \
-pos chr10:117542948 \
-out ./out/NATRIO_chr10_117542948.png \
-read_group strand
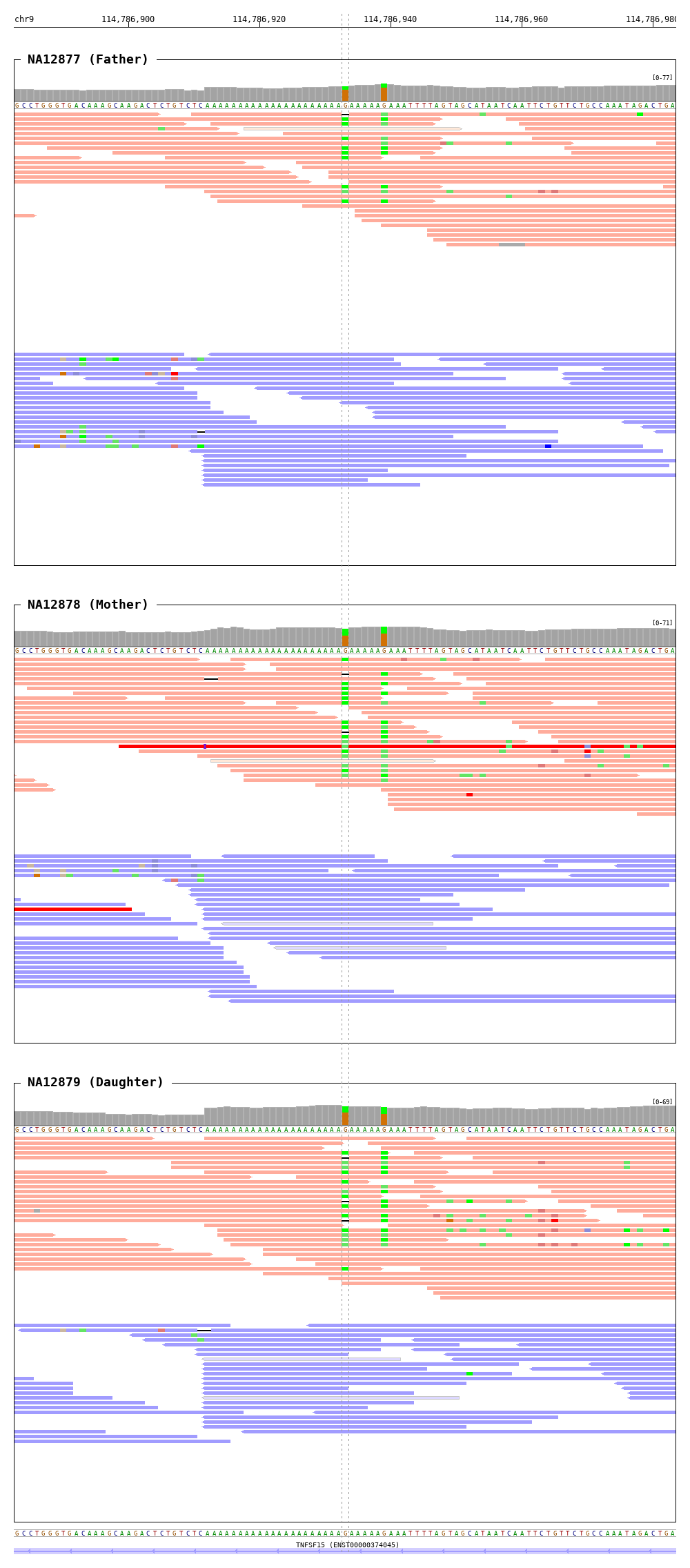
$ bamsnap \
-bam ./data/NA12877.bam \
./data/NA12878.bam \
./data/NA12879.bam \
-title "NA12877 (Father)" "NA12878 (Mother)" "NA12879 (Daughter)" \
-pos chr9:114786933 \
-out ./out/NATRIO_chr9:114786933.png \
-draw coordinates bamplot base gene \
-bamplot coverage base read \
-margin 50 \
-read_group strand \
-plot_margin_left 20 \
-plot_margin_right 20 \
-border

$ bamsnap \
-bam ./data/NA12879.bam \
-pos chr10:117542948 \
-no_title \
-draw bamplot \
-bamplot coverage \
-out ./out/NATRIO_chr10_117542948_3.png \
-separator_height 0
$ bamsnap \
-bam ./data/NA12879.bam \
-pos chr10:117542948 \
-no_title \
-draw coordinates \
-out ./out/NATRIO_chr10_117542948_coordinates1.png \
-no_target_line \
-coordinates_axisloc bottom
Color by inter-chromosomal rearrangements
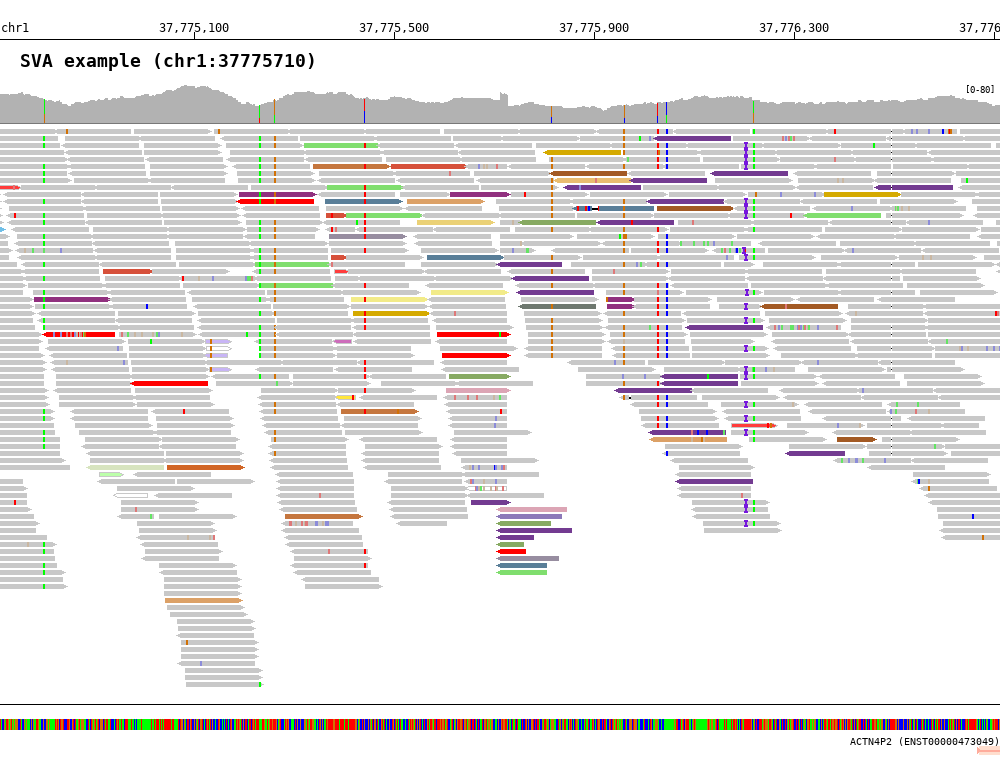
$ bamsnap \
-bam ./data/test_SV1_chr1_37775710.bam \
-title "SVA example (chr1:37775710)" \
-pos chr1:37775710 \
-out ./out/test_SV1-3.png \
-bamplot coverage read \
-margin 1000 \
-no_target_line \
-read_color_by interchrom \
-save_image_only
Deletion
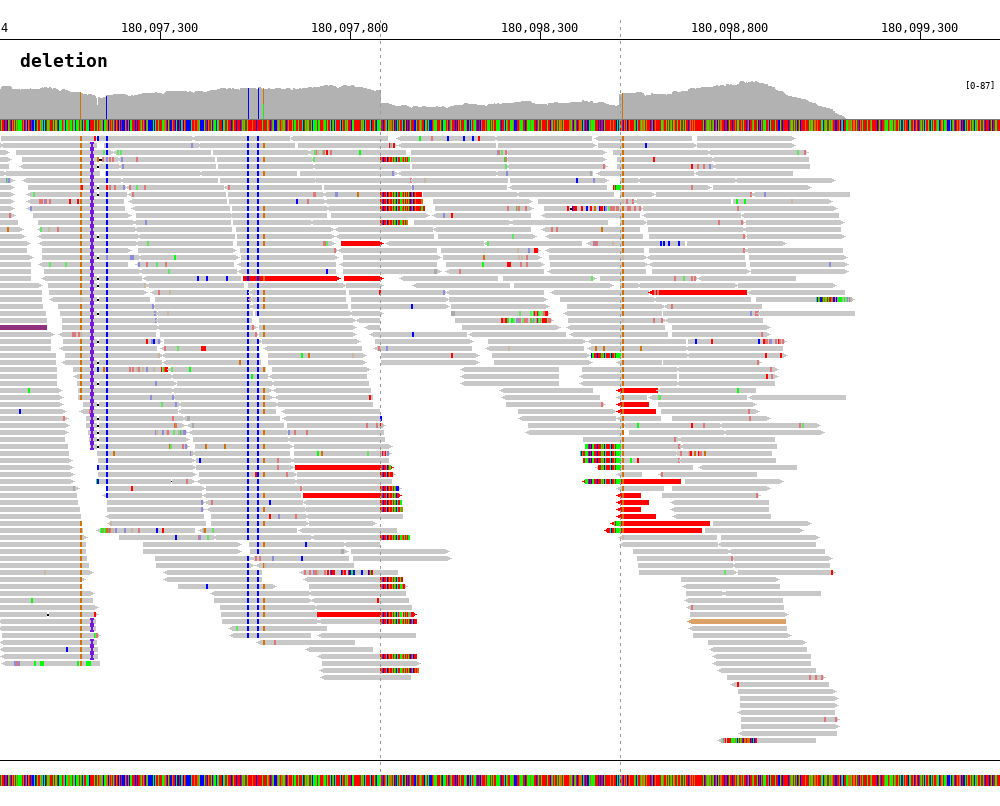
1 2 3 4 5 6 7 8 9 10 | $ bamsnap \
-bam ./data/test_DEL_4_180097876_180097877.bam \
-pos 4:180097878-180098507 \
-margin 1000 \
-title deletion \
-out ./out/test_DEL_1.png \
-refversion hg19 \
-show_soft_clipped \
-read_color_by interchrom \
-save_image_only
|
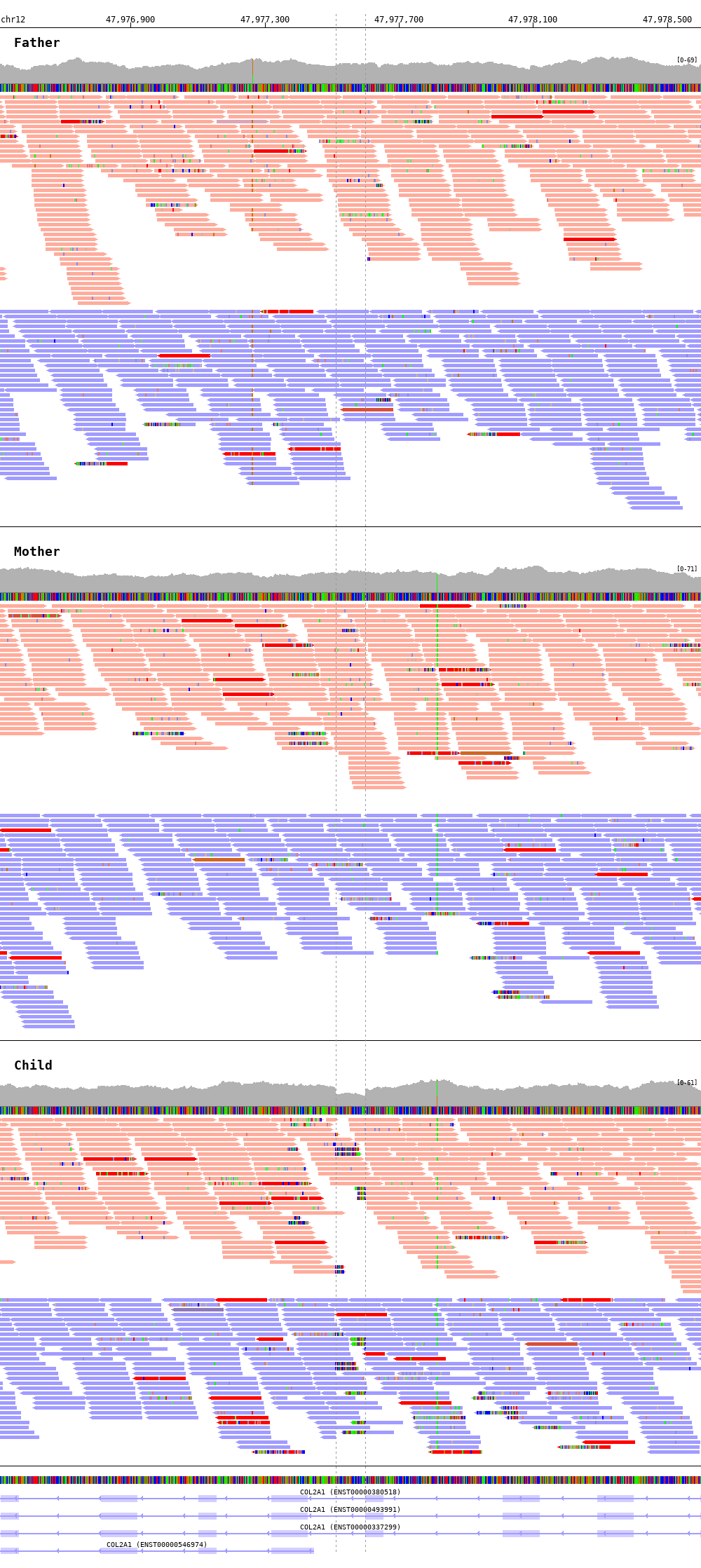
1 2 3 4 5 6 7 8 9 10 | $ bamsnap \
-bam ./data/test_DEL_chr12_47977510_F.bam ./data/test_DEL_chr12_47977510_M.bam ./data/test_DEL_chr12_47977510_P.bam \
-vcf ./data/test_DEL_chr12_47977510.vcf \
-margin 1000 \
-title "Father" "Mother" "Child" \
-out ./out/test_DEL_chr12_2.png \
-show_soft_clipped \
-read_color_by interchrom \
-read_group strand \
-save_image_only
|